Differential Expression AI Exercise
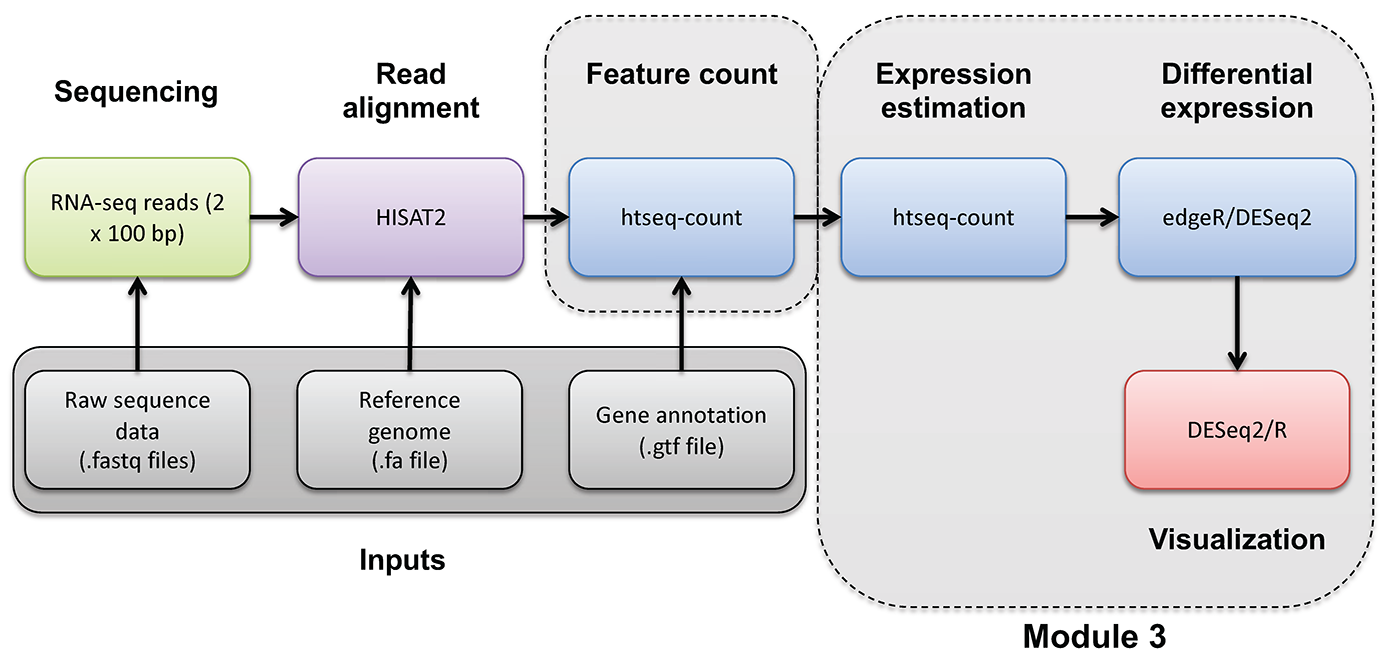
DE Analysis. AI exercise
In this tutorial you will:
- Experiment with use of AI tools to perform a differential expression analysis
- Consider alternative approaches, best practices, pitfalls for use of AI for such work
- Share prompts and compare results with your colleagues
Complete a Google form as you go through this exercise
We will use a google form to capture basic information about AI tool choice (and model version), AI prompts used, etc. Refer to the course slack channel for a link to this form.
Outline of the exercise
For this exercise I want you to imagine that you did not have the previous section as a guide on how to perform a differential expression analysis in R. Imagine that you obtained the gene counts matrix “gene_read_counts_table_all_final.tsv” from your sequencing core, a collaborator, or a public source. See if you can complete a DE analysis like the one you just completed step-by-step but using an AI assistant. Here is an overview of the basic steps (more details on each below):
- Locate the input data and define a location for results
- Choose an AI tool and make note of the model used (record your choice in the Google form)
- Develop your initial prompt to the AI (record your prompt in the Google form)
- With help from the AI create a differential expression analysis in R and run it.
- Answer a few specific questions and produce a visualization for comparison of results to your colleagues.
1. Input data and output location
For the exercise use the following:
- Input gene expression counts data file path:
/cloud/project/data/bulk_rna/gene_read_counts_table_all_final.tsv - Gene ID to name mapping file path:
/cloud/project/data/bulk_rna/ENSG_ID2Name.txt.gz - Results path:
/cloud/project/outdir/de_ai_exercise
Hint. To quickly see what the input data looks like, you can do the following to display the first two lines of each:
head(read.delim("/cloud/project/data/bulk_rna/gene_read_counts_table_all_final.tsv"), n=2)
head(read.delim(gzfile("/cloud/project/data/bulk_rna/ENSG_ID2Name.txt.gz")), n=2)
2. Choice of AI tool
There are several broad approaches to using AI for bioinformatics analysis, and your choice of tool will shape how the interaction unfolds. The main paradigms are:
-
Conversational AI assistants. You interact with the AI through a chat interface, describing your task in plain language and exchanging messages back and forth. You paste in data snippets or describe your files, and the AI generates code or explanations in response. This is the most accessible entry point and requires no special software setup.
-
AI-integrated coding environments. Some development environments embed AI assistance directly into the coding workflow. The AI can see your open files, suggest completions inline, explain errors in context, and generate code without leaving the editor. This tighter integration can make iteration faster but requires a compatible editor.
-
Locally-run open models. Open-weight language models can be run on your own machine, which avoids sending data to external servers (relevant when working with sensitive or unpublished data). These require more setup and capable hardware, and are generally less capable than leading commercial models at time of writing.
Note that given the cloud-based RStudio environment used in this course, the conversational assistant approach is the most practical. Using this approach you can work in a browser tab alongside your RStudio session with no additional setup. The IDE-integrated approach is possible but would require downloading the input data files to your laptop. Locally-run models are beyond the scope of this exercise.
For this exercise, use whichever AI tool you are comfortable with, or take the opportunity to try one you have heard about but not used before, or simply pick one from the list below. A few commonly used options across these categories:
| Tool | Type | Provider |
|---|---|---|
| ChatGPT | Conversational | OpenAI |
| Claude | Conversational | Anthropic |
| Gemini | Conversational | |
| Microsoft Copilot | Conversational | Microsoft |
| GitHub Copilot | IDE-integrated | Microsoft/GitHub |
| Cursor | IDE-integrated | Cursor |
| Ollama | Local models | Open source |
Make note of both the tool and the specific model version you use (e.g. GPT-4o, Claude Sonnet 3.7, Gemini 2.0 Flash). Model versions vary significantly in capability and the field moves quickly. Record your choice in the Google form.
3. Develop initial prompt
Before starting, take a few minutes to think about what you will tell the AI. A well-constructed prompt is likely to produce more useful code than a vague one, and the differences across students will make for a rich comparison later. You do not need to craft a perfect prompt! Part of the point of this exercise is to see what happens with different approaches.
As you draft your prompt, consider what information the AI would need to produce useful, runnable code. Some dimensions to think about (without being told exactly what to include):
- The goal. What kind of analysis do you want? What output would you consider a success?
- The data. What does the input file look like? How is it structured? What do the rows and columns represent? You may want to paste a small excerpt so the AI can see the format directly.
- The experimental design. The AI has no way to know which samples belong to which condition unless you tell it. How would you convey this?
- Tool or method preference. There are several well-established R packages for differential expression analysis. Do you want to specify one, or let the AI choose? Does it matter?
- Filtering and quality considerations. Raw count matrices often contain genes with very low or zero counts across samples. Do you want the AI to handle this, and if so, how?
- Output and thresholds. What criteria define a “differentially expressed” gene for your purposes? What visualizations or result files would be useful?
Write your initial prompt, then record it in the Google form before sending it to the AI. You may refine it through the conversation.
4. Complete the differential expression analysis
Enter your prompt into the AI tool of your choice and examine the response carefully before running anything. Consider:
- Does the response make sense at a high level? Does the AI seem to have understood your goal?
- What assumptions did it make: about the data format, the experimental design, the choice of method, filtering strategy, or significance thresholds? Are those assumptions correct?
- Are there obvious errors or things that seem off?
- Do you feel the initial response is complete enough to run, or do you want to refine it with follow-up prompts before proceeding?
Iterate with the AI as needed. When you are satisfied, copy the code into your Posit cloud environment and execute it line by line. At each step, inspect the output and try to understand what the code is doing before moving on. If something fails or produces unexpected results, you may go back to the AI to debug.
Record your answers to the above questions, along with any significant follow-up prompts, in the Google form.
5. Evaluate the outcome
Before wrapping up, make sure your analysis produces results that address the following. These are the questions you will answer in the Google form and the basis for class discussion:
- Top DE genes. What are the top differentially expressed genes in each direction (upregulated and downregulated)? Report gene names, fold changes, and adjusted p-values.
- Summary statistics. How many genes were tested? How many pass your significance threshold (both in total and broken down by direction)?
- Volcano plot. Produce a volcano plot with the top genes labeled. Make sure it is clean enough to share — axis labels, a title, and clear highlighting of significant genes.
Record your answers in the Google form and save your volcano plot image to your results directory. Be prepared to share it for class discussion. The variation across students in both results and plot appearance will be part of what we examine together.
6. Further reading and exploration
The following resources are starting points for deepening your understanding of the concepts touched on in this exercise. This is a fast-moving area. Treat these as entry points rather than definitive answers.
Comparing and choosing AI tools
- The LMSYS Chatbot Arena leaderboard provides crowd-sourced, head-to-head comparisons of language models across a wide range of tasks and is one of the most widely cited independent benchmarks for conversational AI.
- Most major providers publish their own model capability summaries and release notes. These may be worth checking when new versions are released, as capabilities can change substantially between versions.
Prompt engineering
- The Prompt Engineering Guide (DAIR.AI) is a comprehensive, open community resource covering core techniques.
- Most AI providers also publish their own prompting guides. Anthropic’s guide and OpenAI’s guide are both readable and practical.
- Key concepts to explore further: chain-of-thought prompting, use of examples (zero-shot, one-shot, few-shot, many-shot), role assignment, context engineering (account settings, project config files, system prompts, and other techniques that automatically influence every interaction with the AI), and finally how to iteratively refine prompts based on output quality.
Critical evaluation and responsible use
- Hallucination is the tendency of language models to generate plausible-sounding but factually incorrect information. This may include for example: fabricated function names, wrong default parameters, and non-existent R packages.
- Reproducibility is a growing concern when AI-generated code is used in scientific analysis. The same prompt can produce different code on different days or with different model versions. Document your prompts, the model and version used, and the code that was actually run (not just the final result).
- Data privacy. Commercial AI tools send your input to external servers. For unpublished data, sensitive patient data, or data under a data use agreement, verify the terms of service before sharing anything. Many institutions have policies on this.
- Search for recent perspectives on “LLMs in scientific research reproducibility” and “AI-generated code reliability” for ongoing community discussion of these issues.
AI tools for bioinformatics and data science
- The use of LLMs for bioinformatics tasks such as code generation, literature summarization, and experimental design is an active research area. Search “large language models bioinformatics” in PubMed or Google Scholar for recent reviews and benchmarks.
- Evaluation papers comparing how well models handle statistical analysis, R code generation, and genomics tasks are beginning to appear and are worth tracking.
- Community forums such as Biostars and Bioconductor support forums increasingly include discussions of AI-assisted workflows and how people are actually using these tools.
